PLSC 1.0 Syntax
Draft status: This content reflects the current working spec and may be revised.
1. What is PLSC?
Phenotype List String Code (PLSC) pairs a PL-String with its namespace and version (or date) so that every expression is self-contained and traceable.
<namespace>#<version or date>#<pl-string>
- Namespace — who defines the vocabulary (e.g.,
who,optn,et,nmdp) - Version or date — a release tag (preferred) or ISO date (YYYY-MM-DD)
- PL-String — a phenotype expression using the operators below
Use a release version when available (e.g.,
3.61.0). If not, use the date the reagent/result/interpretation was produced.
2. Namespaces
A namespace represents a governed, versioned code system (or a set of systems with one base). Examples:
who— Base: IPD-IMGT/HLA release; Extensions: WHO antigen names, NMDP MAC codes for ambiguity, and other well-documented, versioned tables.optn— OPTN antigen/epitope codes used in U.S. allocation workflows.et— Eurotransplant antigen/match determinant tables.nmdp— registry codes (including MACs), aligned to specific IMGT/HLA releases.
Gene family namespace
Each gene family namespace represents one or more code systems. When more than one code system is used, a base system is designated which is then extended by the other code systems.
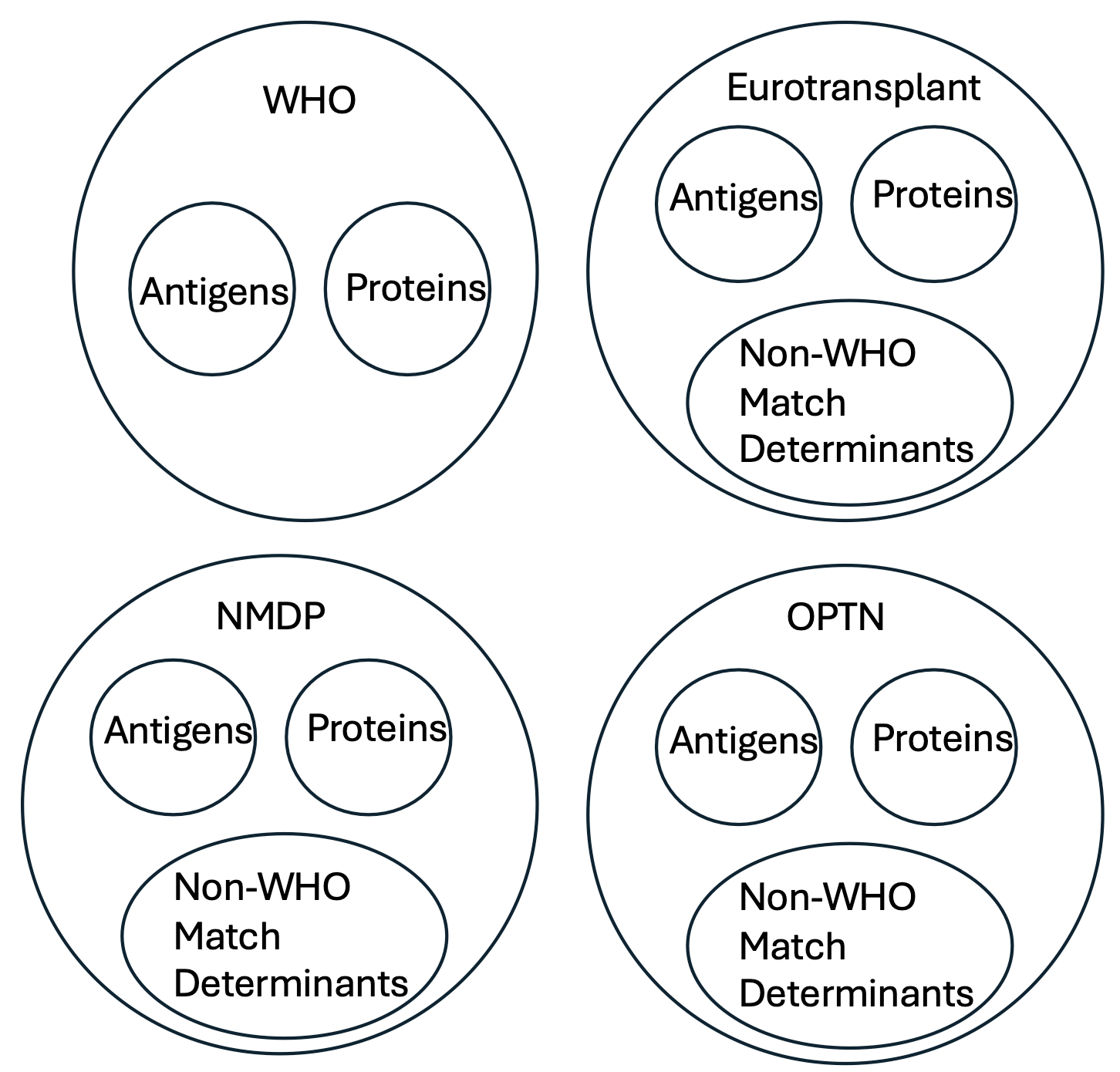 |
- who — Base: IPD-IMGT/HLA release; Extensions: WHO antigen names,
NMDP MAC codes for ambiguity, and other well-documented, versioned tables.
- optn — OPTN antigen/epitope codes used in U.S. allocation workflows.
- et — Eurotransplant antigen/match determinant tables.
- nmdp — registry codes (including MACs), aligned to specific IMGT/HLA releases.
|
Rules
- A PLSC must specify exactly one namespace and one version (or date).
- No mixing of namespaces within a single PL-String.
- Each implementation should publish its governance, release process, and documentation for users.
3. PL-String grammar
PL-String encodes phenotypes (not genotypes). It represents the presence or possible presence of proteins/antigens and heterodimer pairing for class II. It does not carry chromosomal phase, zygosity, or locus order.
3.1 Delimiters and precedence
| Precedence | Delimiter | Name | Meaning (phenotype context) | Typical context |
|---|---|---|---|---|
| 1 | + |
AND | All listed molecules/antigens are present (e.g., on a bead; in a result) | All contexts |
| 2 | ~ |
Heterodimer | Exactly two elements form one heterodimer (class II α~β) | DQ, DP, DR (protein/antigen) |
| 3 | % |
Inclusive OR | One or more of the listed items may be present | Antigen lists; multi-reactivity |
| 4 | / |
Exclusive OR | Exactly one of the listed items is present (ambiguity) | Protein-level ambiguity (e.g., MAC) |
Notes:
+binds most tightly (1),/loosest (4) for unambiguous parsing.%(inclusive OR) is primarily for antigen ambiguity/association./(exclusive OR) encodes protein ambiguity (e.g., unresolved allele).~is only valid for appropriate α~β pairs per the chosen namespace.
3.2 Basic rules
- No loci/phase semantics: PL-Strings do not assert chromosomal phase,
cis/trans, zygosity, or locus order;
DPA1*01:04~DPB1*02:02is equivalent toDPB1*02:02~DPA1*01:04. - Slash locus constraint: Terms on either side of
/must be the same locus/category within the namespace (e.g.,HLA-A*01:01/HLA-A*01:02is valid; mixingAwithBis not). - Heterodimers:
~may only connect permitted α~β pairs as defined by the namespace (e.g.,DQA1~DQB1,DPA1~DPB1,DRA~DRB3/4/5or their antigen equivalents). - No mixed namespaces inside a single PL-String.
- Whitespace is not significant; implementers should normalize.
3.3 Examples
Reagent (protein ambiguity on a bead)
HLA-A*01:01/HLA-A*01:02+HLA-A*02:01
A*02:01is present;A*01:01orA*01:02(but not both) is present.
Result (antigen-level inclusive ambiguity)
A0101%A0102+A2402
A2402 present; A0101 and/or A0102 may be present.
Class II heterodimers (protein level)
HLA-DQA1*05:01~HLA-DQB1*02:01%HLA-DPA1*01:03~HLA-DPB1*04:02
One or both heterodimers may be present.
Constrained heterodimer variants
HLA-DPA1*01:04~HLA-DPB1*02:01%HLA-DPA1*01:04~HLA-DPB1*04:02
One or two heterodimers;
DPA1*01:04is always present.
Multiple loci (phenotype list)
HLA-A*01:02+HLA-A*02:01+HLA-B*07:02
A*01:02,A*02:01, andB*07:02present.
Implementations should reject invalid pairs like
DQA1*05:01~DPB1*04:02based on namespace pairing rules.
4. PLSC construction
Syntax
namespace#version_or_date#plstring
Version: Prefer authoritative release tags (e.g., 3.61.0).
Date: If no release applies, use ISO YYYY-MM-DD. The date reflects when
the reagent/result/interpretation was produced—not sample collection/export.
4.1 Examples
- IPD-IMGT/HLA release bound protein list:
who#3.61.0#HLA-A*01:01/HLA-A*01:02+HLA-A*24:02 - Class II heterodimers with antigen mapping:
who#2025-11-02#DQA1*05:01~DQB1*02:01%DPA1*01:03~DPB1*04:02 - Ambiguous MAC usage (namespace-governed):
nmdp#2025-11-02#HLA-DQB1*03:01/297+HLA-DQB1*06:02
5. Validation checklist
- Namespace present and recognized.
- Version or date present and well-formed.
- No mixed namespaces within
<pl-string>. - Operators parse with precedence:
+>~>%>/. - Slash locus constraint holds on every
/. - Heterodimer pairs are permitted by the namespace.
- Atoms exist in the declared namespace release/date.
6. Embedding in exchange formats
- FHIR: Use as a
Codingwithsystem(e.g.,http://plstring.org),version(namespace release or date), andcode(the full PLSC string). - HAML: Store as the code value for reagent/result/interpretation elements.
- No mixing of namespaces inside one code value; use separate codings for different namespaces.
FHIR example
<valueCodeableConcept>
<coding>
<system value="http://plstring.org"/>
<version value="1.0"/>
<code value="who#3.61.0#HLA-DQA1*05:01~HLA-DQB1*02:01+HLA-DPB1*04:01"/>
</coding>
</valueCodeableConcept>
7. Comparison to GL-String (at a glance)
| Concept | GL-String (genotype) | PL-String (phenotype) |
|---|---|---|
| Focus | Alleles/genotypes & phase | Proteins/antigens & heterodimers |
| Locus delimiter | ^, |, ? (various) |
Not used |
| Exclusive OR | / (allele ambiguity) |
/ at protein level |
| Inclusive OR | (N/A) | % at antigen level |
| Heterodimer | ~ for phase |
~ for α~β pairing (class II) |
| Composition | + |
+ (reagent contents; multi-reactivity) |
| Mix namespaces | Allowed per implementer | Disallowed within a single PL-String |
8. Frequently asked
Can I encode epitopes/eplets?
Not in this version. Once stable, widely adopted nomenclatures are available,
the grammar can extend to epitope atoms (e.g., OPTN DPB1 unacceptable
epitopes, eplet registries).
What about non-classical loci (E, F, G) or MICA/MICB/ABO?
Permitted if your namespace publishes governed vocabularies; apply the same
operators and rules.
This page supersedes earlier drafts and aligns operator semantics (+, ~,
%, /), constraints (no mixed namespaces; heterodimer pairing rules), and
PLSC binding (namespace + version/date) with the current manuscript. It replaces
the older GL-centric placeholder text.